Sparse encoding and finding solutions of the form Ax=b with x subject to an l1 norm is a popular solution in signal reconstruction and now face recognition. Unlike the least-squares l2 norm which can be solved rapidly with SVD, finding the sparse l1 norm is a rather CPU expensive task. One of the fastest l1 solvers, GPSR still takes several seconds per image to process. If you profile the code, it turns out that 95% of the time is spent in matrix multiplication, which got me to thinking “there must be more efficient ways to do this.” I came up with three ways to optimize GPSR:
1) Single precision floating point math
Matlab by default uses double precision floating point math, so one quick and dirty solution is to convert everything to singles before calling GPSR. This gives a speed up of roughly 50%, which is fairly effective with a very negligible drop in accuracy (<0.5%) because of the lower precision.
2) Batch processing with matrices
The way I use GPSR is to find the sparse representation of one test face from a set of training faces. It is not unusual to have tens of thousands of training images, each with a size of 100-300 after PCA dimensionality reduction. Say we have 25,000 training images each with dimensionality of 100. Using GPSR, the bottleneck in finding the sparse representation of a test image involves multiplying a 25,000×100 matrix by a 100 vector many times until some convergence (I am of course leaving out some other things that happen for simplicities’ sake). By matrix multiplication properties, we can process multiple test images in batch by concatenating their vectors into a 100xN matrix. Although accomplishing the same thing, this single matrix multiply has the advantage of a single BLAS call where things can remain in L1 and L2 cache longer, giving us a much higher speedup. The problem here is some test images converge faster than others, but we can get around this by dynamically inserting and removing test image vectors into our concatenated “test images” matrix as needed. This yields about a 5X improvement with a batch size of 256, bringing us to about a 10X improvement over the original GPSR – for this particular application, of course.
3) GPU acceleration
A final step to optimizing the GPSR algorithm is to move to specialized hardware, such as graphic cards via a CUDA interface. Luckily, nVidia provides a CUDA implementation of BLAS called cuBLAS. How much speed ups could would we be likely to attain using this? Since matrix multiplication is still roughly 50% of my optimized GPSR algorithm, I took a look at matrix multiplication on various architectures.
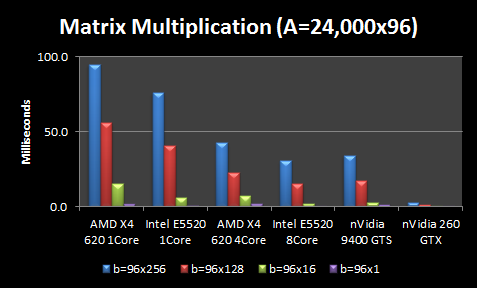
I conducted two test configurations with some simplifying assumptions. First, I assumed all matrix sizes are multiples of 16 because it seems cuBLAS seems to prefer this memory configuration. So I assume our training matrix A is 24,000×96 and test images are 96×1. I consider both without batching and with batches of test images with various sizes: 16, 128, and 256. Thus, I vary the test images matrix b from sizes ranging from 96×1 to 96×256.
All tests are averaged over 10-100 matrix multiplications with single precision matrices. I tested 4 CPU configurations and 2 GPU configurations:
- AMD X4 620 1Core ($100): Low-end 2.6 GHz Athlon quadcore system without L3 cache with only 1 core used
- AMD X4 620 2Core ($100): Low-end 2.6 GHz Athlon quadcore system without L3 cache with all 4 cores used
- Intel E5520 1 Core ($375): High-end 2.26 Xeon quadcore system with only 1 core used
- Intel E5520 2 Core($750): Two high-end 2.26 Xeon quadcores with all 8 cores used
- nVidia 9400 GT ($40): Low-end GPU with only 16 CUDA cores
- nVidia 260 GTX ($200): Middle-end GPU with 216 CUDA cores
NOTE: CPU configurations used MATLAB, which in turn usually uses CPU-optimized BLAS libraries like MKL on Intel or ACML on AMD. For a quick comparison, I benchmarked the the fast GotoBLAS against Matlab with the MKL and found GotoBLAS only 2% faster. Since this performance increase is fairly negligible, I didn’t pursue testing different BLAS packages.
NOTE: Memory transfer is not included in these comparisons because I can leave the large A matrix on the GPU unmodified as I process thousands of test images. Thus my problem is floating-point bound rather than memory bound.
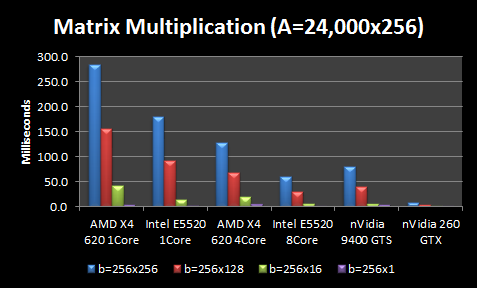
I also consider the same scenario except the image size is 256 instead of 96 (so more PCA dimensions).
The CPU results are not terribly surprising. More expensive CPUs outperform less expensive CPUs, but the performance gains for more expensive CPUs are not proportional to the extra cost (i.e. 4X increase in CPU cost doesn’t give you anywhere close to a 4X performance boost). Throwing more cores at the problem helps, but again the scaling is nowhere near perfect. An extra 4 cores gives you ~2X boost and 8 cores gives you ~3X boost in performance. Larger matrices see better scaling. The interesting part is the GPUs. A $40 nVidia GPU is better than the $100 AMD CPU using all 4 cores, and performs about equivalently if you factor in the memory transfer to the GPU.
In fact, the $40 GPU is only beat by my $750 dual Intel CPUs with 8 cores, which is kind of astonishing. And the middle-end $200 nVidia GPU outperforms everything by a wide margin, averaging 5-30X better. However, at these high processing speeds, data transfer becomes the overhead. Thus, the Intel 8 Core still beats the nVidia 260 GTX if we factor in memory transfer of the large A matrix for every multiplication. However, I only have to transfer the large A matrix of ~25,000 training images to the GPU once. Thus for my application, it could still be tremendously faster to use the GPU.
Although it doesn’t show well on this graph, using matrix multiplication for b=96×1 is actually 1.5X slower than b=96×16 , leading me to believe that GPUs do have some interesting side-effects when it comes to matrix sizes, block optimizations, etc. It is good to know these intricacies because you could get significant speedups just by zero-padding matrices so the dimensions are divisible by 16.
I haven’t actually coded up the GPSR algorithm in CUDA simply because my existing 10X speedups seem to be working pretty well so far. And if I run GPSR on multiple datasets simultaneously, one on each core, I should be able to get much better parallelism than distributing each matrix multiplication over many cores. However, it is interesting to know that a $40 graphics card and some CUDA programming could outperform a brand new quad-core machine.